
Dean’s Quarterly
The Icahn School of Medicine at Mount Sinai is a national and international leader in biomedical education, research, and patient care.
Dean’s Quarterly
The Dean's Quarterly features the latest news and research from Icahn School of Medicine at Mount Sinai.
This report highlights:
- A Message from the Dean: 2018 has been another strong year for the Icahn School of Medicine at Mount Sinai
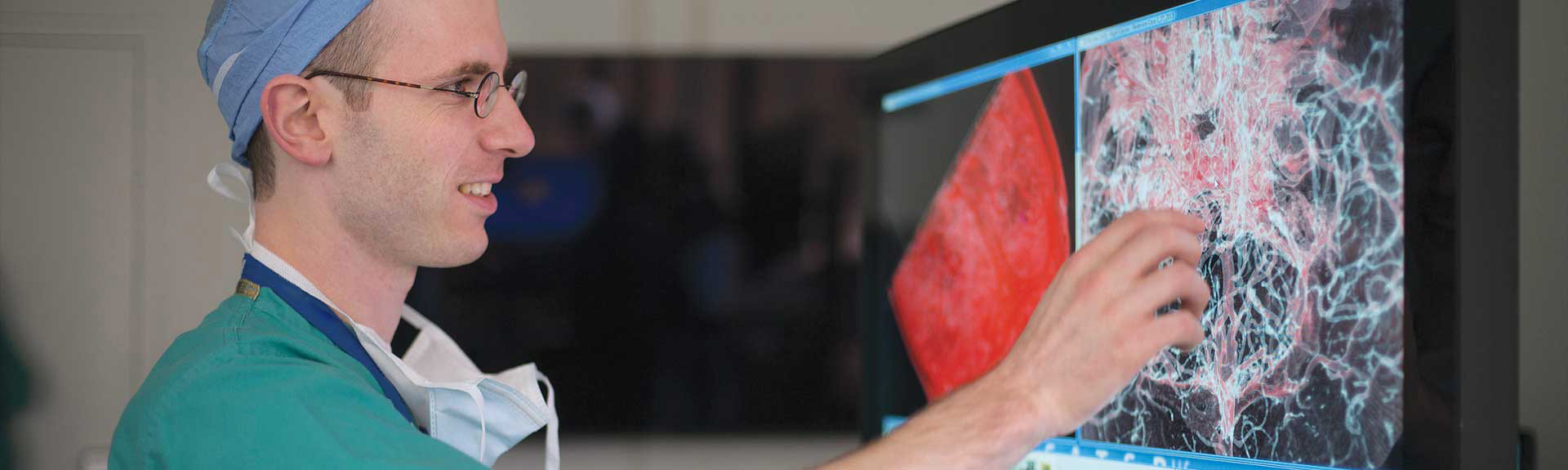
- Researchers using artificial intelligence to detect acute neurological illness in minutes
- A data-driven study that identifies the role of human herpesvirus in Alzheimer's disease
- Computer models for a new era of precision medicine
- Reprogramming retina cells to restore vision that may lead to treatment for severe degenerative eye disease
- Improving the prognosis for melanoma patients
- A computational model that leads to discovery of gene-activation pathway associated with atherosclerosis
- New studies that provide insight into p53-like bladder cancer that is generally resistant to chemotherapy
- Mount Sinai becoming a key player in emerging blockchain technology
- A novel RNA/DNA sequencing platform that may help determine the best drugs for certain multiple myeloma patients
- Two studies that point to the quality of neoantigens in long-term cancer survival
Message from the Dean

This has been a strong year for the Icahn School of Medicine at Mount Sinai. Our National Institutes of Health (NIH) funding for 2018 increased by 9.4 percent over 2017, reaching $348.5 million—the highest level in our 50-year history—and placing us at No. 12 among all U.S. medical schools. Our basic science departments in genetics, microbiology, neuroscience, and pharmacological sciences were among the top four in NIH funding, and we ranked fourth in research dollars per principal investigator among U.S. medical schools.
Mount Sinai’s robust residency program continued to be the largest in the United States in 2018, with more than 2,500 residents and clinical fellows. With a footprint that extends throughout the Greater New York area, Mount Sinai offers trainees and house staff unprecedented opportunities to care for one of the nation’s most diverse population of patients.
The students in our medical school, MD/PhD program, PhD program, and master’s programs also embody a spectrum of talents and accomplishments. Recently, we launched our Biomedical Science Initiative, which will integrate the work being done in computing and big data analytics across Mount Sinai and support our goal of becoming a premier institution for genomic and data science.
The Mount Sinai Health System's new capital campaign is underway with a goal of raising $1.5 billion. This campaign will enable us to leverage our size, along with our clinical and research strength, in areas that include oncology, cardiology, immunology, and drug discovery. We plan to establish centers of excellence in neurotherapeutics, precision medicine, immunotherapeutics, and genome editing; hire additional faculty; build a new life science center; and invest in genomic medicine and predictive analytics. We will continue to foster partnerships with industry and create new Mount Sinai companies based on our intellectual property.
We expect our new campaign to be as transformational as our last one was 10 years ago, when we began planting the seeds for discovery that are now bearing fruit. Mount Sinai’s investments in supercomputing, bioinformatics, and advanced technologies have led to many of the novel findings that have been published in top scientific journals. You will read about several of these studies in the following pages of Dean's Quarterly.
Dennis S. Charney, MD, Anne and Joel Ehrenkranz Dean, Icahn School of Medicine at Mount Sinai, and President for Academic Affairs, Mount Sinai Health System
Artificial Intelligence Detects Acute Neurological Illness in Minutes
Mount Sinai Health System researchers have designed an automated deep-learning neural network that can detect acute neurological diseases such as stroke, hemorrhage, and hydrocephalus within minutes, enabling earlier intervention that can preserve neurological function and improve patient outcomes.
This is the first study to use artificial intelligence to identify acute neurological disease and thus demonstrate a direct clinical application in a randomized controlled trial. Using 37,236 head T scans, researchers trained a 3D conventional neural network (3D-CNN) to determine whether an image contained critical or noncritical findings, thus enabling triaging and queuing of radiology workflow for physician review based on the probability of acute neurological illness. Researchers subsequently conducted a randomized, double-blinded prospective trial to assess the ability and quickness of the automated deep-learning neural network to recognize and provide notification of a critical finding compared to that of two radiologists and a neurosurgeon in a simulated clinical environment.
The results of the study, which were published August 2018, in Nature Medicine, revealed that the 3D-CNN was able to process, interpret, and alert physicians to a critical finding in 1.2 seconds, compared to an average human time of three minutes. Researchers also noted a significant difference in the queue position of urgent versus routine cases in the 3D-CNN and human cohorts, with acute cases appearing earlier in the automated deep-learning neural network queue.
“This is a system with the capacity to raise alarms about critical findings in mere seconds and thus prevent these scans from potentially sitting in a queue for minutes, even hours, before they are assessed by a specialist,” says the study’s lead author, Eric K. Oermann, MD, an Instructor of Neurosurgery at the Icahn School of Medicine at Mount Sinai. “Our trial demonstrated that using artificial intelligence to identify cases of acute neurological illness works, and for that reason we should embrace this technology.”
“We know that rapid response is critical in the treatment of acute neurological illnesses,” says study co-author Joshua B. Bederson, MD, Professor and Chair of the Department of Neurosurgery, Mount Sinai Health System, and Clinical Director of the Neurosurgery Simulation Core. “Any tools that decrease time to diagnosis may lead to improved patient outcomes.”
The study was conducted using “weakly supervised learning approaches,” which resulted in a lower diagnostic precision for the 3D-CNN platform compared to physicians (0.48 vs. 0.85) at a comparable sensitivity (0.79). Researchers are now focusing on re-engineering the platform using “strongly supervised learning approaches” and novel techniques that will enhance diagnosis accuracy. The expectation is that this work will be completed within the next two years.
“The application of deep learning and computer vision techniques to radiological imaging is a clear imperative for twenty-first century medical care,” says study author Burton Drayer, MD, the Charles M. and Marilyn Newman Professor and Chair of the Department of Radiology for the Mount Sinai Health System, and Dean for Clinical Affairs, Icahn School of Medicine at Mount Sinai.
This study was performed by the Mount Sinai AI Consortium (AISINAI), a group of scientists, physicians, and researchers dedicated to developing artificial intelligence platforms that enhance clinical diagnosis and patient care. AISINAI’s research builds on a paper published in January 2018 in Radiology that details the consortium’s work on natural language processing algorithms capable of identifying clinical concepts in radiology reports for CT scans.
Data-Driven Study Identifies the Role of Human Herpesvirus in Alzheimer's Disease
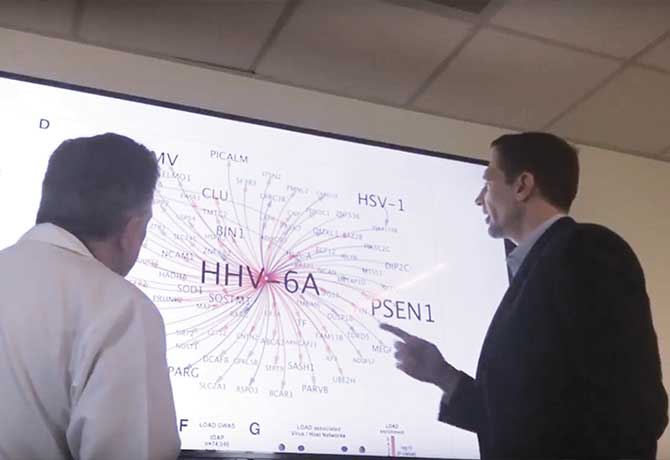

Two strains of human herpesvirus—human herpesvirus 6A (HHV-6A) and human herpesvirus 7 (HHV-7)—are found in the brains of individuals with Alzheimer’s disease at levels up to twice as high as in those without Alzheimer’s, according to findings published June 21, 2018, in Neuron by a team of Icahn School of Medicine at Mount Sinai researchers. These common viruses, best known for causing roseola, a skin rash in children, can also be present in the brain and remain inactive for years. They also can cause encephalitis and other chronic disorders.
It is the first study to use an entirely data-driven approach to examine the impact of viruses on Alzheimer’s—and to identify potential disease-associated roles of these particular viruses, which was an unexpected discovery. The researchers had been comparing genetic data in healthy and postmortem Alzheimer’s brains to identify possible new drug targets for Alzheimer’s disease when their complex computations revealed the unusually high levels of the viral genomes.
“This study represents a significant leap forward in our understanding of how viruses may play a role in Alzheimer’s disease,” says the study’s senior author, Joel Dudley, PhD, Mount Sinai Professor in Biomedical Data Science, and the Director of the Icahn School of Medicine’s Institute for Next Generation Healthcare. “We were able to not only measure these viruses using computational techniques but also build out the networks in which these viruses are operating and influencing known Alzheimer’s genes.”
The study was enabled in part by powerful new molecular profiling data released by the National Institute on Aging (NIA) Accelerating Medicines Partnership-Alzheimer's Disease, a collaboration among industry, government, and nonprofit organizations dedicated to identifying new targets for preventing or treating the disease. Using those data, the team
conducted computer mapping of four brain regions from more than 600 samples from the Mount Sinai Brain Bank, which for the last 36 years has been examining postmortem brain tissue from donors for Alzheimer’s-related research. As the study progressed, the team found the higher levels of HHV-6A and HHV-7 viruses, which was subsequently confirmed using data from brain banks in three other major NIA Alzheimer’s Centers. Researchers also discovered that the viruses appeared to accelerate the development of Alzheimer’s-related plaques and tangles in the brain.
“Our team was initially quite skeptical about these findings,” says the study co-senior author, Sam Gandy, MD, PhD, Mount Sinai Professor in Alzheimer’s Research and the Director of the Center for Cognitive Health and NFL Neurological Care at Mount Sinai. “Gradually, we became convinced that what we were seeing is a real property of Alzheimer’s brains in several internationally recognized brain banks, and that we ought to report it, even if we cannot completely explain what these viruses are doing in the brain.”
Although the study does not prove a direct causal role between these herpesviruses and Alzheimer’s disease, or indicate what activates the viruses, the findings do lend credence to the hypothesis that these herpesviruses may trigger progression of dementia in Alzheimer’s. This work also opens new avenues for the identification of subpopulations and biomarkers, which would not only make it easier to diagnose and determine a person’s risk of developing Alzheimer’s but also possibly create opportunities to use existing antiviral drugs and drugs that stop the brain’s immune cells from responding to these viruses.
“We have been inundated with letters and emails from people sharing stories about viral infection and Alzheimer’s disease in their personal or family life, and that is an impetus for us to find an effective therapy as soon as we can,” Dr. Dudley says.
Developing Computer Models for A New Era of Precision Medicine

Mount Sinai’s Institute for Data Science and Genomic Technology and three outside institutions have partnered to create a new center that will advance precision medicine by developing comprehensive predictive models of biological systems.
With $6.5 million from the National Institutes of Health, the new Center for Reproducible Biomedical Modeling is a collaborative effort among Mount Sinai; the University of Washington; the University of Connecticut Health Center; and the University of Auckland, in New Zealand.
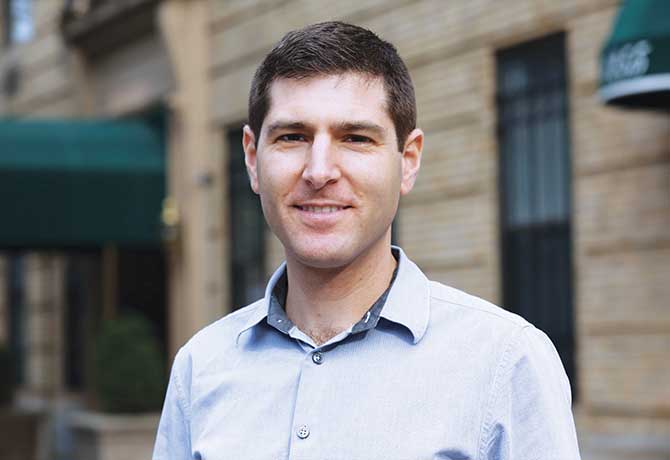
Advancing precision medicine and bioengineering requires an integrated understanding of how biological behavior arises from the millions of genes and genetic interactions in cells. “Mechanistic models, particularly models of cells, are needed to help us understand this enormous complexity,” says Jonathan Karr, PhD, Assistant Professor, Department of Genetics and Genomic Sciences, and a Project Director of the Center.
“Many engineering fields routinely use mechanistic models to design complex systems such as planes. In contrast, medicine and bioengineering are often slow, ineffective, and expensive because we lack similar models of biology. Our goal is to help physicians precisely treat disease by developing comprehensive mechanistic models of whole cells,” Dr. Karr says. “Whole-cell models are one of the grand challenges of the twenty-first century. Building whole-cell models will require substantial technological innovation. The Center will develop some of the first technologies for scalably modeling and simulating entire cells.”
Developing the technology needed to build comprehensive models of cells will provide scientists with a better understanding of the molecular basis of behavior and enable bioengineers to design microorganisms that sense and disrupt disease.
Building comprehensive, predictive models will also require extensive interdisciplinary collaboration and huge investment. To build the team needed to achieve whole-cell models, the Center will organize conferences, seminars, tutorials, and challenges.
“Ultimately, we envision that physicians will treat disease by sequencing each patient’s genome and using mechanistic models to identify the best therapy, or combination of therapies, for each patient,” Dr. Karr explains.
To bring that vision to reality, Dr. Karr, along with Arthur Goldberg, PhD, Associate Professor, Department of Genetics and Genomic Sciences, and their collaborators are pioneering comprehensive models of cells and developing the technologies needed to enable these models.
Dr. Karr says, “Armed with the technologies developed by the Center, we expect to prove that these sophisticated models are possible, and that they will transform how we practice medicine.”
To learn more about the Center and follow its latest research, visit the Center’s website at https://reproduciblebiomodels.org.
Reprogramming Retina Cells to Restore Vision May Lead to Treatment for Severe Degenerative Eye Disease
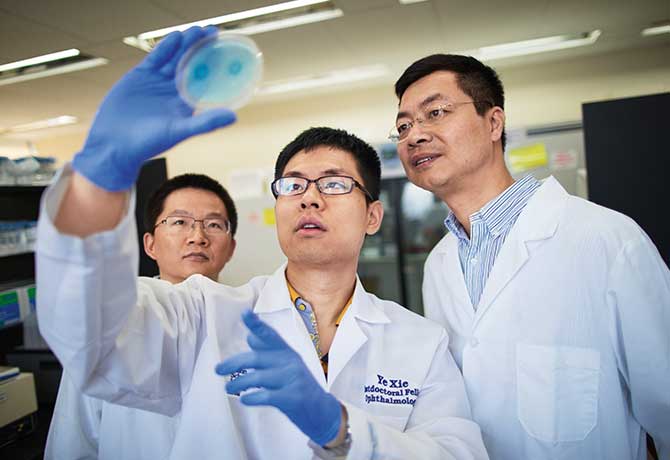

A team of researchers, led by Bo Chen, PhD, Associate Professor of Ophthalmology, Icahn School of Medicine at Mount Sinai, has reversed congenital blindness in mice by changing the supportive Müller glia cells in the retina into functional rod photoreceptors—light-sensitive cells in the retina that signal the brain when activated.
The breakthrough research, published online in the August 15, 2018, issue of Nature, is expected to advance efforts toward retinal regeneration for diseases of the eye, such as age-related macular degeneration, degenerative glaucoma, and retinitis pigmentosa. Funded in part by the National Eye Institute (NEI), an arm of the National Institutes of Health, the study drew praise from NEI Program Director Thomas N. Greenwell, PhD.
“This is the first report of scientists reprogramming Müller glia to become functional rod photoreceptors in the mammalian retina,” says Dr. Greenwell. “Rods allow us to see in low light, but they may also help preserve cone photoreceptors, which are important for color vision and high-visual acuity. Cones tend to die in later-stage eye diseases. If rods can be regenerated from inside the eye, this might be a strategy for treating diseases of the eye that affect photoreceptors.”
Scientists have long studied the regenerative potential of Müller glia cells because in species such as zebrafish, they divide in response to injury and can turn into photoreceptors and other retinal neurons. In the lab, scientists have coaxed mammalian Müller glia to behave as they do in the fish, but not without injuring the tissue. Since injured tissue is counterproductive to restoring vision, Dr. Chen’s lab was able to achieve an effective method of achieving both goals, something others could not.
In pursuing this novel gene-transfer therapy, Dr. Chen, who is also Director of the Ocular Stem Cell Program at the Icahn School of Medicine at Mount Sinai, and his investigators followed a two-stage process. First, they demonstrated that Müller glia cells could be spurred to divide in mice by injecting their eyes with a gene to turn on a protein called beta-catenin. Then, weeks later, the mice were injected with transcription factors Otx2, Crx, and Nrl that encouraged the newly divided cells to develop into rod photoreceptors. The investigators were encouraged to find that the newly formed cells looked structurally no different from real photoreceptors, and that synaptic structures that allow the rods to communicate with other types of neurons within the retina had also formed.
When the researchers took another step forward by testing the treatment in congenitally blind mice that were born without functional rod photoreceptors, the results were positive once again. The light responses recorded from retinal ganglion cells—neurons that carry signals from photoreceptors to the brain—and measurements of brain activity confirmed that the newly formed rods were integrating into the visual pathway circuitry, from the retina to the primary visual cortex in the brain.
“Our findings underscore that we are closer than ever to developing new therapies for people with severe degenerative eye disease,” says Dr. Chen. “Mice that were blind from birth were now able to see light for the first time following treatment.” Dr. Chen says his next step will be determining whether the technique works on cultured human retinal tissue.
Improving the Prognosis for Melanoma Patients
A novel epigenetic mechanism that causes resistance to the standard treatment given to melanoma patients with mutations in the BRAF gene has been discovered by Emily Bernstein, PhD, and her team of researchers at the Icahn School of Medicine at Mount Sinai. BRAF mutations are found in roughly half of all melanomas. The researchers also found a biomarker—a protein called IGFBP2—that is associated with poor prognosis in melanoma patients.
Both findings appear in the August 24, 2018, issue of Nature Communications, and point to a potential multipronged approach to preventing drug resistance or reversing it once it has occurred, by inhibiting the effects of BRAF mutations and IGFBP2-driven biological pathways.

“The incidence of cutaneous malignant melanoma is rising and its therapeutic management remains challenging,” says Dr. Bernstein, an Associate Professor of Oncological Sciences and Dermatology. “In recent years, there has been extensive therapeutic development to inhibit key biological targets. Although a large proportion of patients with advanced metastatic melanoma harboring BRAF mutations respond to the standard therapy, known collectively as MAPK inhibitors, subsequent resistance remains a major clinical challenge.”
Dr. Bernstein is conducting further studies on the epigenetic mechanisms altered in cancer, including the modeling of macroH2A knockout mice models for melanoma and breast cancer.
“The H2A family is the largest family of histone variants and it has been largely understudied in cancer,” Dr. Bernstein says. “We know histone variants can promote tumorigenic programs, so we want to target these proteins and prevent them from doing that. But we cannot target the histone on its own because there is no aspect of its structure where we could apply a small molecule that would block its role.”
Instead, Dr. Bernstein is looking at potentially targeting the “histone chaperone-histone variant” interaction that introduces these proteins into the genome. “We are setting up assays and working together with the Drug Discovery Institute at the Icahn School of Medicine to identify small molecules that could block this interaction,” she says. “We are still in the very early stages, but if this research is successful, it opens the door to collaborating with medicinal chemists on an enhanced molecule that can be tested in animals.”
According to Dr. Bernstein, the more researchers delve into the epigenome, the more targets they will discover. “There are many startups and large pharmaceutical companies already moving into this line of drug development and some treatments are already undergoing clinical trials,” Dr. Bernstein adds. “It is clear that alteration of the epigenome is a major driver in multiple cancers and the more we explore that, the better we will understand how it works.”
Computational Model Leads to Discovery of Gene-Activation Pathway Associated with Atherosclerosis

A computational model of cells that line blood vessels in the human heart, developed at the Icahn School of Medicine at Mount Sinai, has led to the discovery of a gene-activation pathway caused by lipids associated with coronary artery disease. The findings appear in the June 12, 2018, issue of Nature Communications.
The new pathway, discovered by researchers in Mount Sinai’s Department of Genetics and Genomic Sciences, and at Goethe University in Frankfurt, Germany, could help identify new directions in research and drug development.
Atherosclerosis is caused by the buildup of a complex mixture of components, commonly referred to as plaque, within the inner lining of arteries. Oxidized phospholipids are abundant in this arterial plaque and are thought to promote atherosclerosis progression. However, the specific cellular processes caused by these lipids on the arterial surface are still not well understood. The cells composing the inner surface of blood vessels, called endothelial cells, are at the forefront of the atherosclerotic process and, therefore, are a major focus of research into coronary artery disease.
“Computational biological models such as the one we used in this study are allowing us to uncover a wealth of knowledge about complex diseases that we never could before,” says Jun Zhu, PhD, Professor of Genetics and Genomic Sciences at the Icahn School of Medicine at Mount Sinai; Head of Data Science at Sema4, a patient-centered predictive health company that is a Mount Sinai venture; and co-senior author of the study. “Endothelial cell response to lipids has been studied extensively over the years, but it was still unknown that MTHFD2 was even functional in these cells.”
The researchers from Mount Sinai and Goethe University predicted and validated in follow-up experiments that the MTHFD2 gene plays a key role in endothelial cell response to oxidized phospholipids. They found that MTHFD2 was also activated in endothelial cells in response to other factors, such as inflammation or a change in amino acid concentration. This underscores the many factors involved in the development of atherosclerosis that must be understood and taken into consideration when approaching disease therapies.
“Our study showed that when the MTHFD2 gene is activated in endothelial cells in response to oxidized lipids, it sends out molecular ‘danger signals’ promoting inflammation and stimulating the atherosclerotic process,” says Ralf Brandes, MD, Director of the Institute for Cardiovascular Physiology and Professor of Physiology at Goethe University. “These findings suggest that MTHFD2 could be a novel target to disrupt development and progression of atherosclerosis.”
While the role of MTHFD2 in the vascular system was unknown before this study, the gene is known to be consistently activated in cancer, making it a promising target for cancer therapies. In clinical trials, MTHFD2 inhibitors are already in use as anti-cancer therapies.
“It’s possible that these therapies could also help prevent coronary artery disease, but more research into the specific role of MTHFD2 in atherosclerosis is needed first, before proposing it as a target for potential therapy,” according to Dr. Zhu.
New Studies Provide Insight into p53-like Bladder Cancer That Is Generally Resistant to Chemotherapy
Two microRNA biomarkers that can help stratify which patients with aggressive p53-like bladder cancer will have a better or worse prognosis have been uncovered by researchers at the Icahn School of Medicine at Mount Sinai. Until now, the optimal approach to treating p-53-like tumors—which are generally resistant to chemotherapy—has been poorly defined and urgently needed.
“Working with our colleagues in Genetics and Genomic Sciences, we applied a computer model that was able to make microRNA much more understandable in terms of its impact on tumor behavior,” says Matthew Galsky, MD, Professor of Medicine ( Hematology and Medical Oncology) at The Tisch Cancer Institute at the Icahn School of Medicine. That research—published in the July 2018 issue of Oncogene— identified two specific microRNAs as potential prognostic markers of the p53-like subset that might also have the ability to yield “critical information for personalizing treatment options,” he adds.

The study’s computer model, known as ActMIR, had previously been developed at Mount Sinai and successfully validated microRNA activity in multiple subtypes of breast cancer. Researchers now applied it to bladder cancer genomic data in the National Cancer Institute’s Cancer Genomic Atlas to identify the activities of the two biomarkers of p53-like bladder cancers—miR-106b-5p and miR-532-3p.
“Our computational methods not only provided us with deeper insights into the cellular mechanisms of this elusive type of bladder cancer, but revealed the potential of microRNAs as therapeutic targets for treating the disease,” says the study’s senior author, Jun Zhu, PhD, Professor of Genetics and Genomic Sciences at Mount Sinai, and Head of Data Science at Sema4, a Mount Sinai venture.
Additional discoveries about the sensitivity and resistance of bladder cancer to immunotherapy were published by the research team in the August 2018 issue of Nature Communications.
The team’s goal is to bring their findings to the clinic and establish early-stage clinical trials that will improve patient care. Almost two years ago, The Tisch Cancer Institute established the Novel Therapeutics Unit in order to accomplish this. To date, more than 15 clinical trials are under way at the Novel Therapeutics Unit, some involving new drugs and others involving novel combinations of existing drugs.
“A critical mission of any cancer center is to be able to offer patients cutting-edge therapies that can provide the best clinical outcomes,” says Dr. Galsky, Director of the Novel Therapeutics Unit. “That’s why our approach through the Novel Therapeutics Unit is to capitalize on today’s most promising science originating from within the walls of Mount Sinai, as well as outside from our industry partners."
Mount Sinai Becomes a Key Player In Emerging Blockchain Technology
As blockchain moves from the world of cryptocurrency to the field of health care, the Mount Sinai Health System is poised to take advantage of this potentially transformative technology. In July, Mount Sinai created the Center for Biomedical Blockchain Research to examine how blockchain might provide effective solutions to problems affecting health care and the biomedical sciences. The Center will also build and test its own blockchain systems.
“Blockchain builds on our expertise in biomedical data, machine learning, and data governance, and could, ultimately, help us improve health care delivery and reduce costs,” says Noah Zimmerman, PhD, Assistant Professor of Genetics and Genomic Sciences at the Icahn School of Medicine at Mount Sinai, and Director of the Center for Biomedical Blockchain Research. “The technology is still in its very early stages, but we’re determined to be ahead of the curve as the infrastructure gets built and people figure out where the most promising applications lie.”
Blockchain entered the public domain some 10 years ago as the transactional engine behind Bitcoin. In brief, blockchain is like a digital ledger or spreadsheet where all information is shared with other members as part of a large, collaborative, and decentralized network. “Blockchain, at its core, is about building large-scale open networks and incentivizing people to use them,” explains Dr. Zimmerman. “In the case of health care, blockchain technology could provide exciting new ways to store, access, and process data.”
Underscoring its potential is the fact that approximately 160 companies have raised more than $660 million for health care-related blockchain projects over the past three years. Investors and health care experts see the most promising applications in areas that include giving patients greater access to and control over their personal medical records, promoting greater collaboration among health care institutions, distributing and sharing clinical trials data, and engaging patients to seek treatment, take their medicines, and participate in clinical research.
The Center for Biomedical Blockchain Research has been tracking blockchain initiatives in health care as part of its public database. While conducting scholarly evaluations of blockchain-enabled solutions, the Center also plans to form partnerships with companies looking to develop blockchain initiatives in clinical medicine and biomedical research. Dr. Zimmerman reports that there is interest, and one joint venture has already been launched with an early stage digital health start-up.
“There’s a lot of excitement around the possibilities of blockchain technology in health care,” acknowledges Joel Dudley, PhD, Associate Professor of Genetics and Genomic Sciences at Mount Sinai and Director of the Institute for Next Generation Healthcare, which houses the Center for Biomedical Blockchain Research.
Citing “the fragmented nature of regional and global health care systems,” Dr. Dudley envisions blockchain as a way to overcome current obstacles to the flow of vital information that denies access to underserved groups. “Blockchain and related technologies could potentially enable applications that result in more unified health care ecosystems,” he says, “and thus serve the greater goals of realizing national and global precision health networks.”
A Novel RNA/DNA Sequencing Platform May Help Determine the Best Drugs for Certain Multiple Myeloma Patients

As multiple myeloma progresses and patients exhaust multiple lines of therapy, the options for treatment become fewer. Relapse is nearly inevitable and median survival stands at around six years. Researchers at Mount Sinai have taken steps to alter those odds by developing a novel RNA and DNA sequencing platform for determining which drugs work best for late-stage and drug-resistant multiple myeloma patients.
In a clinical trial reported in JCO Precision Oncology in August 2018, they expanded the traditional DNA-based approach by adding RNA sequencing to uncover targets for a broad swath of drugs approved by the U.S. Food and Drug Administration that go well beyond those typically used to treat multiple myeloma, the second most common blood cancer. The work of these researchers represents the first time genome-wide RNA profiles have been clinically assessed for hematological malignancies.
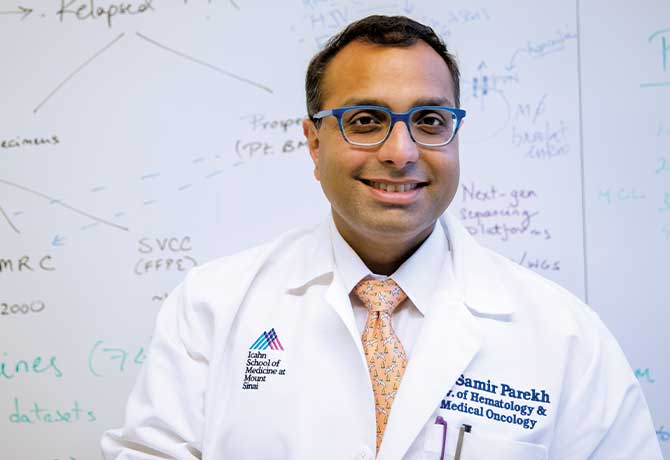
“Our findings really expand the scope of what patients with advanced multiple myeloma can receive,” says Samir Parekh, MD, Associate Professor of Medicine (Hematology and Medical Oncology), and Director of Translational Research in Myeloma at The Tisch Cancer Institute at the Icahn School of Medicine.
“We identified treatment options for each patient by checking different libraries with hundreds of compounds each until we found the best therapeutic drugs.” And of the evaluable trial participants who received an FDA-approved drug based on a comprehensive profile generated from DNA sequencing and RNA data, 76 percent showed a clinical benefit—a response rate Dr. Parekh called “a real eye-opener, especially since many of these patients were end-stage myeloma.” Indeed, the therapy served as a life-saving bridge for some patients to new drugs in clinical development, such as CAR T-Cell, an exciting new immunotherapy that genetically reengineers the body’s T cells to recognize and attack cancer cells in the patient’s bloodstream.
RNA sequencing is viewed by developers as a perfect complement to the move to precision medicine in cancer therapy, where treatment is based on specific genomic alterations observed in a patient’s tumor, rather than on the tumor histology or tissue type. “It’s able to capture more dynamic aspects of unique tumor biology and provide information beyond what’s available with DNA alone,” notes Joel Dudley, PhD, Associate Professor of Genetics and Genomic Sciences at the Icahn School of Medicine, who collaborated with Dr. Parekh on the RNA/DNA profiling study.
Mount Sinai is home to one of the largest multiple myeloma programs in the nation, providing translational research and end-to-end care to more than 400 new patients each year who are often referred by physicians from around the world.
“Because of our patient volume, we’ve developed a unique understanding of myeloma biology and how to look outside the myeloma toolbox to match DNA mutations and RNA profiles to specific drugs,” says Dr. Parekh.
Two Studies Point to the Quality of Neoantigens in Long-term Cancer Survival
The quality, not quantity, of tumor neoantigens may best predict a patient’s response to cancer immunotherapy and his or her chance of long-term survival.
That finding was based on two groundbreaking studies co-authored by Benjamin Greenbaum, PhD, Icahn School of Medicine at Mount Sinai. Now, it is being further explored by Dr. Greenbaum and Vinod Balachandran, MD, Memorial Sloan Kettering Cancer Center, through a Stand Up to Cancer Convergence 2.0 Grant and funding from the Lustgarten Foundation.

Their goal is to understand the underpinnings of pancreatic cancer survivorship, and their team is one of seven research groups that comprise Stand Up to Cancer’s broad $11 million initiative that was announced in January 2018.
“Previous research has shown that T cell immunity is linked to exceptional outcomes for the few long-term survivors of pancreatic cancer, but it wasn’t clear if neoantigens played a role,” says Dr. Greenbaum. “Our newest research shows there are particular neoantigens that seem to be driving this long-term response in patients with pancreatic cancer, and further suggests that targeting those neoantigens might be a viable therapeutic strategy.”
According to Dr. Greenbaum, who was trained as a physicist and quantitative biologist, “One of the most exciting aspects of our work is collaborating with academic teams that can combine the tools and knowledge from diverse fields like theoretical physics, structural biology, mathematics, and computer science, and translational genomics. We never lose sight of the fact that our combined efforts could one day result in clinical breakthroughs that benefit countless numbers of people.”
Earlier research formed the basis of the team’s efforts. In the first of two back-to-back studies published in Nature (November 2017) researchers described a mathematical model they developed—the first of its kind—to predict how a cancer patient would benefit from certain immunotherapies. Dr. Greenbaum was the senior author on this work with first author Marta Luksza, PhD, Assistant Professor of Oncological Sciences, Icahn School of Medicine. By capturing aspects of a tumor’s evolution and ways in which it interacted with the underlying immune system, the new model appeared to offer an approach beyond previous biomarkers. Going forward, it also has the potential to uncover new therapeutic targets within the immune system and help in the design of vaccines for patients who do not respond to immunotherapy.
The second study, whose first author was Dr. Balachandran, applied the modeling framework to better understand immune response in patients with pancreatic cancer and, more specifically, the unique role of neoantigens.
As part of the broad initiative, the Mount Sinai researchers are focused on better understanding what makes a neoantigen a good immune target and the role of the microbiome in neoantigen recognition. By advancing the core science behind developing a vaccine for pancreatic cancer, the research could eventually improve treatment prospects for patients with this deadly form of the disease.
As they investigate the immune system response to cancers, the Convergence 2.0 team will be encouraged to draw upon the knowledge of Microsoft Research experts in machine learning and artificial intelligence. In addition to Dr. Luksza, the research team includes Nina Bhardwaj, MD, PhD, from the Icahn School of Medicine at Mount Sinai; and Eileen M. O’Reilly, MD; Taha Merghoub, PhD; and Jedd D. Wolchok, MD, PhD, from Memorial Sloan Kettering Cancer Center.